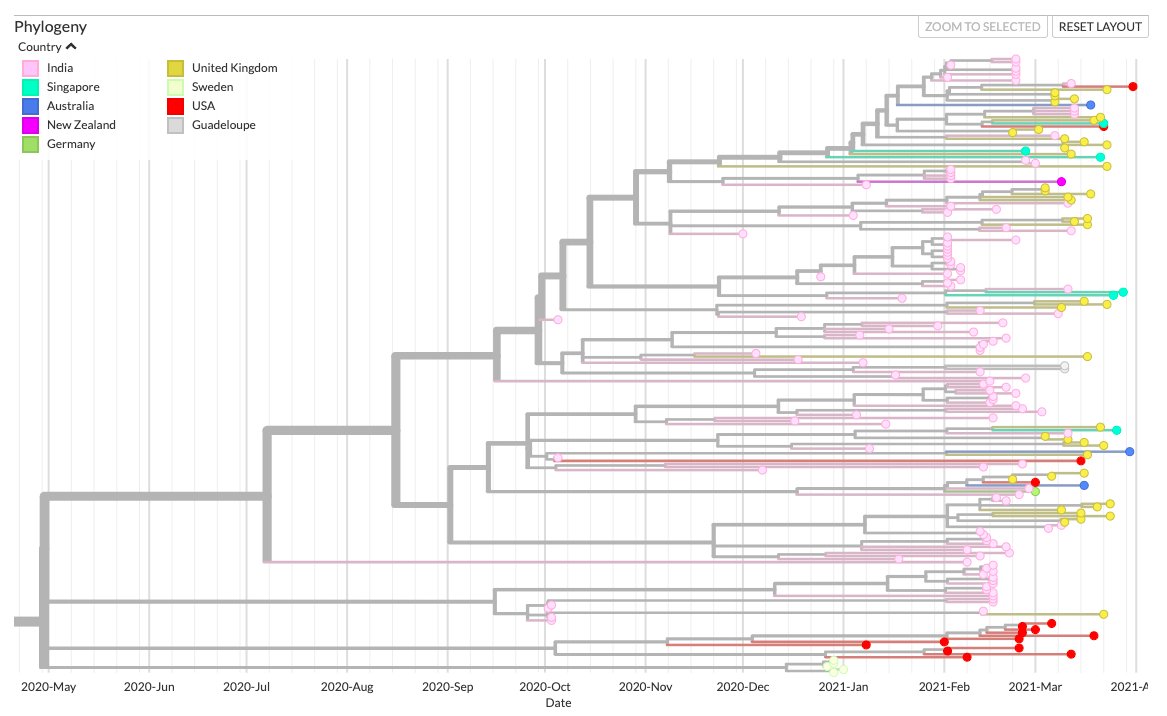
A relatively new emergent #SARSCoV2 lineage/variant from India, bearing the combination of Spike mutations L452R+E484Q, has been making rounds in the news recently. Here& #39;s a quick phylogenetic analysis of all genomes with L452R+E484Q from GISAID.
https://nextstrain.org/community/banijolly/Global-Analysis/484Q-452R?branchLabel=none&c=country
1/7">https://nextstrain.org/community...
https://nextstrain.org/community/banijolly/Global-Analysis/484Q-452R?branchLabel=none&c=country
1/7">https://nextstrain.org/community...
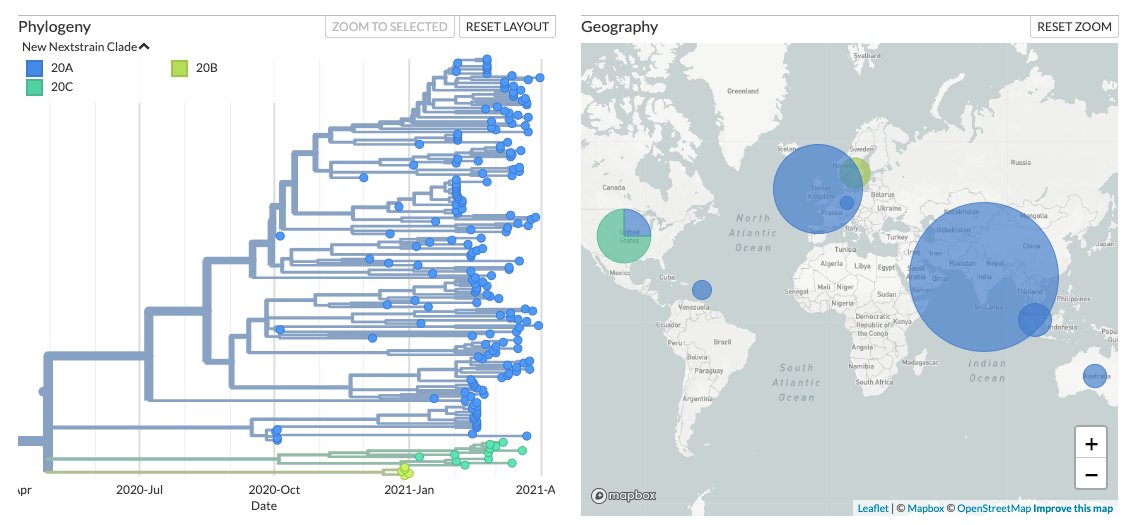
L452R+E484Q seems to be recurrently emerging in 3 different lineages. While the Indian genomes fall under Nextstrain clade 20A, the USA and Sweden genomes form separate clusters under 20C and 20B respectively
@trvrb @nextstrain
https://nextstrain.org/community/banijolly/Global-Analysis/484Q-452R?branchLabel=none&d=tree,frequencies,map&p=grid
2/7">https://nextstrain.org/community...
@trvrb @nextstrain
https://nextstrain.org/community/banijolly/Global-Analysis/484Q-452R?branchLabel=none&d=tree,frequencies,map&p=grid
2/7">https://nextstrain.org/community...
In USA, E484Q was found in some California VOC genomes (PANGO Lineages B.1.427-B.1.429), which already contain L452R
https://twitter.com/firefoxx66/status/1351224782176579584
A">https://twitter.com/firefoxx6... zoomed-in view of the USA B.1.427-B.1.429 cluster:
3/7
https://twitter.com/firefoxx66/status/1351224782176579584
A">https://twitter.com/firefoxx6... zoomed-in view of the USA B.1.427-B.1.429 cluster:
3/7
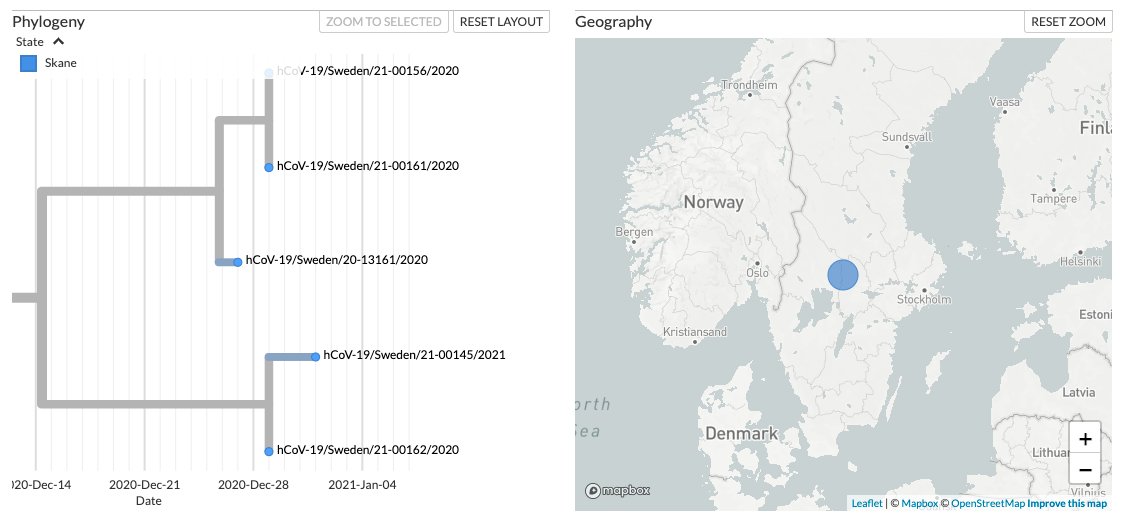
While in Sweden, L452R+E484Q was found in genomes belonging to the lineage L.3 and was detected as early as December 2020 (EPI_ISL_766714)
A zoomed-in view of the Sweden L.3 cluster:
4/7
A zoomed-in view of the Sweden L.3 cluster:
4/7
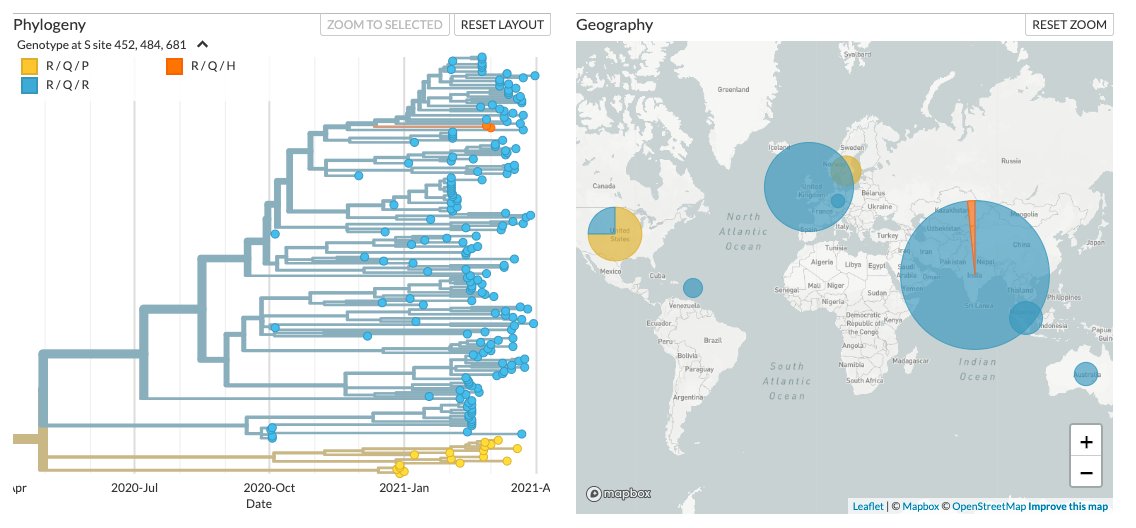
In India, as already previously reported, the variant has shown a particularly increased prevalence in Maharashtra, where the variant is seen to been present at a very low frequency since October 2020
https://nextstrain.org/community/banijolly/Phylovis/COVID-India?branchLabel=none&c=gt-S_452,484&d=tree,frequencies,map&dmax=2021-02-19&dmin=2020-09-08&f_state=Maharashtra&p=grid
5/7">https://nextstrain.org/community...
https://nextstrain.org/community/banijolly/Phylovis/COVID-India?branchLabel=none&c=gt-S_452,484&d=tree,frequencies,map&dmax=2021-02-19&dmin=2020-09-08&f_state=Maharashtra&p=grid
5/7">https://nextstrain.org/community...
And the variant has recently been designated as the PANGO Lineage B.1.617
@arambaut @AineToole
https://cov-lineages.org/lineages/lineage_B.1.617.html
Apart">https://cov-lineages.org/lineages/... from L452R+E484Q, the genomes also have the spike mutation P681R.
6/7
@arambaut @AineToole
https://cov-lineages.org/lineages/lineage_B.1.617.html
Apart">https://cov-lineages.org/lineages/... from L452R+E484Q, the genomes also have the spike mutation P681R.
6/7
Both mutations are associated with antibody escape while L452R has also been associated with a modest increase in infectivity https://covariants.org/variants/S.L452Rhttps://twitter.com/vinodscaria/status/1374778715339509766
Moving">https://covariants.org/variants/... forward, further sequencing and experimental assays will be needed to establish how concerning the combination is!
7/7
Moving">https://covariants.org/variants/... forward, further sequencing and experimental assays will be needed to establish how concerning the combination is!
7/7
 Read on Twitter
Read on Twitter