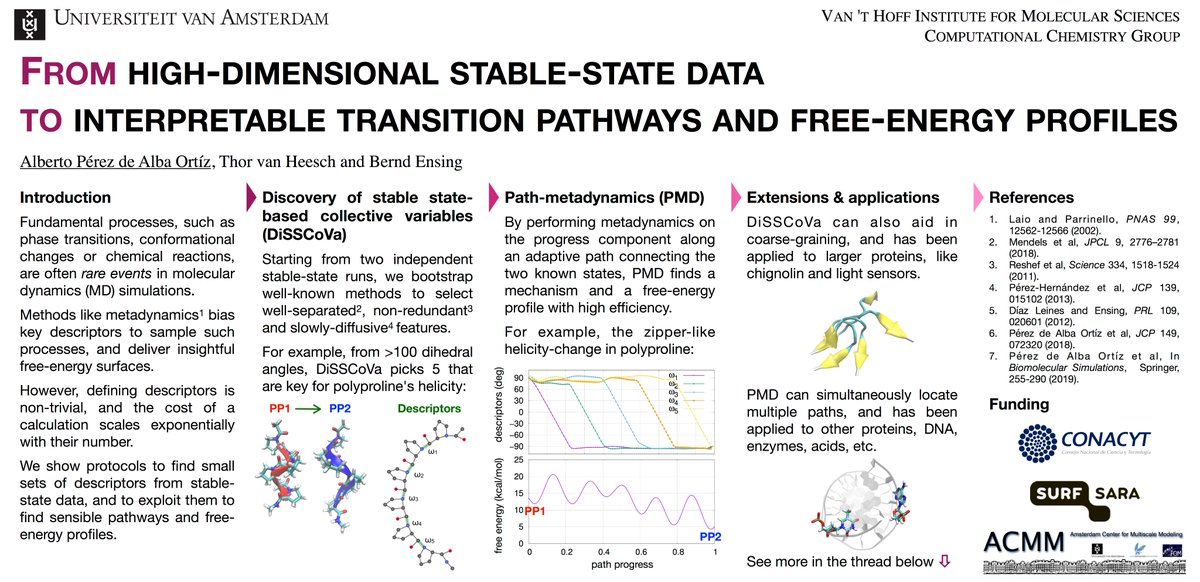
Hi, @LatinXChem! This is my work with @thorvanheesch & @BerndEnsing on data mining & path-based enhanced sampling. Our goal is to go "From high-dimensional stable-state data to interpretable transition pathways and free-energy profiles" #LatinXChem #LatinXChemTheo #Theo154 (1/7).
If you& #39;re curious about #DiSSCoVa, you can watch this one-minute video from the recent @ai4science_lab workshop: https://www.youtube.com/watch?v=KI3BsW7234A">https://www.youtube.com/watch... (2/7)
Here, you can see our favorite didactic animation of #pathmetadynamics in action: https://youtu.be/YRZ3zOO5e_I .">https://youtu.be/YRZ3zOO5e... Notice how the method adapts to find the minimum free energy path! (3/7)
And here, you can see the multiple-path version: https://youtu.be/RNjRAOmQEcw .">https://youtu.be/RNjRAOmQE... Notice how both paths are captured simultaneously! The green path has an slightly higher barrier, but could still compete with the purple one (4/7).
In this blog post, you can read about our application of #pathmetadynamics to base-pairing transitions in #DNA together with Jocelyne Vreede: https://protocolsmethods.springernature.com/posts/51977-pathfinding-in-free-energy-landscapes-of-simple-and-complex-molecular-transitions?channel_id=1911-behind-the-paper.">https://protocolsmethods.springernature.com/posts/519... It will lead you to our book chapter with all you need to know about starting your own runs (5/7).
And talking about starting your own #pathmetadynamics runs, here& #39;s our @PlumedN contribution with a ready-to-run example: https://www.plumed-nest.org/eggs/19/033/ .">https://www.plumed-nest.org/eggs/19/0... You can use it with any MD package supported by @plumed_org! (6/7)
Finally, some further references:u2028
- The original #pathmetadynamics paper: https://doi.org/10.1103/PhysRevLett.109.020601u2028
-">https://doi.org/10.1103/P... Our paper with applications to different systems in classical, ab initio and QM/MM MD: https://doi.org/10.1063/1.5027392
u2028Thanks">https://doi.org/10.1063/1... for reading :) (7/7)
- The original #pathmetadynamics paper: https://doi.org/10.1103/PhysRevLett.109.020601u2028
-">https://doi.org/10.1103/P... Our paper with applications to different systems in classical, ab initio and QM/MM MD: https://doi.org/10.1063/1.5027392
u2028Thanks">https://doi.org/10.1063/1... for reading :) (7/7)
 Read on Twitter
Read on Twitter