Teamwork... This ought to rattle some cages...
1) RaTG13 is incapable of binding to a bat or a pangolin (no binding observed), and is less than 1/1000 the affinity to humans than CoV2 and less than 1/300 the affinity than SARS. It is incapable of forming an infection in humans—
1) RaTG13 is incapable of binding to a bat or a pangolin (no binding observed), and is less than 1/1000 the affinity to humans than CoV2 and less than 1/300 the affinity than SARS. It is incapable of forming an infection in humans—
2) most logical explanation is that RaTG13 is CoV2 with the Spike RBD swapped out in a desperate attempt by Shi to distance herself and her lab from the origin. The RBM itself is in an orphaned Amplicon and a TSS amplicon, both not touching any of the S1 inserts,
3) which is likely the product of an attenyation experiment.
Two: the proposed O-linked glycans downstream of the FCS, do not exist. https://academic.oup.com/glycob/article/doi/10.1093/glycob/cwaa042/5826952
This">https://academic.oup.com/glycob/ar... difference, that between software prediction and experimental result,
Two: the proposed O-linked glycans downstream of the FCS, do not exist. https://academic.oup.com/glycob/article/doi/10.1093/glycob/cwaa042/5826952
This">https://academic.oup.com/glycob/ar... difference, that between software prediction and experimental result,
4) demonstrate a classical example of glycoengineering failure. They attempt to do tissue culture passage and glycoengineering to make it more deadly, including the S1 Insert
5) that is only found in the very low quality and possibly synthetic/snatched amplicons of RaTG13, are all examples of Epitope tagging and are hallmarks of rational protein design. The dead miner virus must have a far more similar RBM in order to begin infection—
6) supported by the RaTG13 RBM being “lost by recombination” (e.g. overwritten on a keyboard). https://europepmc.org/article/PPR/PPR153377
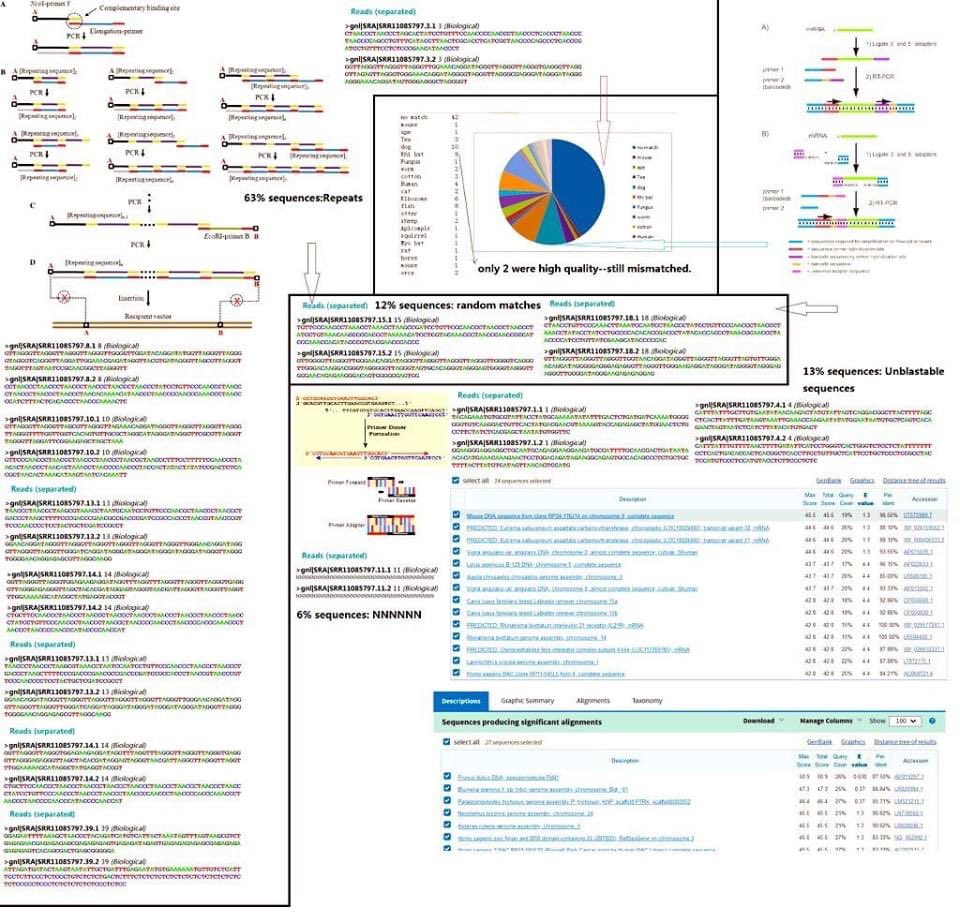
The">https://europepmc.org/article/P... “swab” sequence of RaTG13 displays the hallmark of a PCR reaction on a sample with little to no RNA
The">https://europepmc.org/article/P... “swab” sequence of RaTG13 displays the hallmark of a PCR reaction on a sample with little to no RNA
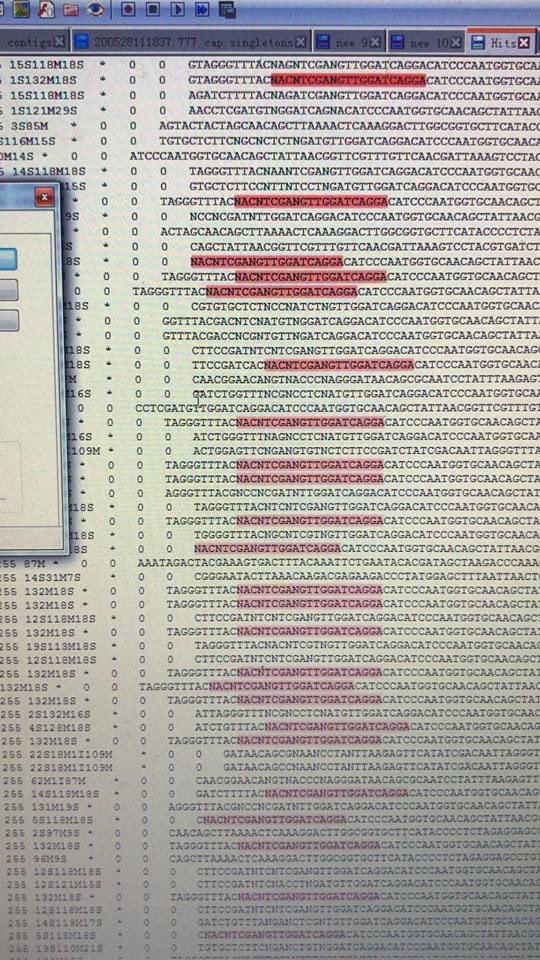
7) —massive amplification of repeats, generation and overpopulation of meaningless random adhesion products, and massive duplication of whatever reads still recognizable there.
Corresponding with a lack of uniquely attributable bacterial reads
Corresponding with a lack of uniquely attributable bacterial reads
8) —the only ones that “matches” to a bacterial genome, matches at the conserved parts of the 35S rRNA that can be either bacterial or mitochondrial. This is inconsistent with a “fecal swab” that should be mostly bacteria. This implies a lack of usable template molecules—
9) meaning that it will be impossible for the mNGS to miraculously cover ths entire genome—most likely it was Spiked with amplicons.
https://zenodo.org/record/3885333#.Xw86COT92Ef
This">https://zenodo.org/record/38... also invalidates the Lam et. al. datasets (below last images)
https://zenodo.org/record/3885333#.Xw86COT92Ef
This">https://zenodo.org/record/38... also invalidates the Lam et. al. datasets (below last images)
10) —as they too show syndromes of intentional read cendorship and filtration—skewing and excluding any possible metagenomic analysis which imply dishonesty
—most likely Spiked with synthetic DNA and filtered.
—most likely Spiked with synthetic DNA and filtered.
@Baric_Lab @PeterDaszak @ChrisRugby7s @BillyBostickson H/T @flavinkins @POTUS @EcoHealthNYC @gatesfoundation
#WeKnow #NotNarural #SARSCoV2 #CCP
#WeKnow #NotNarural #SARSCoV2 #CCP
12) Article which may help all understand the post ;) https://www.minervanett.no/angus-dalgleish-birger-sorensen-coronavirus/the-evidence-which-suggests-that-this-is-no-naturally-evolved-virus/362529">https://www.minervanett.no/angus-dal...
2A) https://twitter.com/flavinkins/status/1283469517901840395?s=21">https://twitter.com/flavinkin... https://twitter.com/flavinkins/status/1283469517901840395">https://twitter.com/flavinkin...
@scrippsresearch #KristianAndersen Not so fast  https://abs.twimg.com/emoji/v2/... draggable="false" alt="😂" title="Gesicht mit Freudentränen" aria-label="Emoji: Gesicht mit Freudentränen">
https://abs.twimg.com/emoji/v2/... draggable="false" alt="😂" title="Gesicht mit Freudentränen" aria-label="Emoji: Gesicht mit Freudentränen">
https://twitter.com/flavinkins/status/1295604749505409026?s=21">https://twitter.com/flavinkin... https://twitter.com/flavinkins/status/1295604749505409026">https://twitter.com/flavinkin...
Human shit! https://twitter.com/flavinkins/status/1295605463547965440?s=21">https://twitter.com/flavinkin... https://twitter.com/flavinkins/status/1295605463547965440">https://twitter.com/flavinkin...
https://twitter.com/flavinkins/status/1295606068639219713?s=21">https://twitter.com/flavinkin... https://twitter.com/flavinkins/status/1295606068639219713">https://twitter.com/flavinkin...
 Read on Twitter
Read on Twitter