In this final thread of my #LipidTakeover, I want to showcase some of the new and exciting research in the field of mass spectrometry for lipid analysis that aims to overcome the limitations of the technique.
Please note this list is not exhaustive, there are many great people in the field doing excellent work and I apologise in advance if I forget anyone! Please feel free to respond to this thread highlighting work in this field that I might have missed
So you& #39;ve already heard me talk about the work of Steve Blanksby and Todd Mitchell, and the development of both OzID and RDD for lipid structure elucidation. Specifically, OzID can be used for determining double bond position and stereochemistry, as well as regiochemistry
But plenty of other researchers out there are also using ozone for lipids structure research, either in the back-end of mass spectrometers or being introduced to lipids prior to ionisation.
In particular, I want to highlight the work of Qibin Zhang at @UNCG who first implemented OzID on a travelling wave ion mobility cell (Waters Synapt G2). https://dx.doi.org/10.1002/rcm.7920">https://dx.doi.org/10.1002/r...
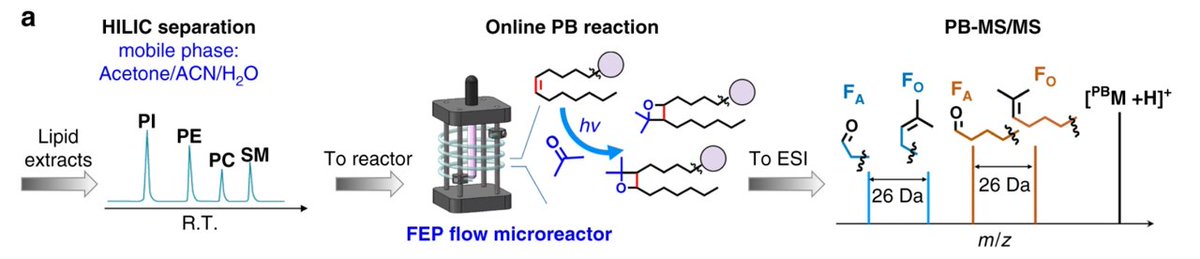
Another excellent technique for C=C position detection is that of Yu Xia from @PurdueChemistry and @Tsinghua_Uni that uses the Paternò-Büchi reaction in order to pinpoint the location of double bonds in a wide range of lipid classes.
Most recently, Yu Xia and colleagues have implemented this workflow on a large scale to determine C=C position in more than 200 unsaturated glycerophospholipids http://www.nature.com/articles/s41467-018-07963-8">https://www.nature.com/articles/...
Photodissociation is also a useful technique for lipids structure elucidation. @gavinreid of @unimelb uses 193 nm ultraviolet photodissociation to discover C=C position in sphingolipids and other lipid species https://dx.doi.org/10.1007/s13361-017-1668-1">https://dx.doi.org/10.1007/s...
Likewise, Jennifer Brodbelt at @UTChemistry also uses UVPD@193 nm to detect structural features in a range of bacterial lipids http://dx.doi.org/10.1021/jacs.7b06416">https://dx.doi.org/10.1021/j...
Takeshi Baba at @sciex has developed electron impact excitation of ions from organics (EIEIO) to successfully determine regiochemistry as well as C=C position and stereochemistry and in a wide range of lipid classes https://doii.org/10.1021/acs.analchem.5b01460">https://doii.org/10.1021/a...
(Side note, Steve Blanksby managed to convince Takashi to use the phrase "old macdonald had a farm" in his talk at ASMS one year - either 2016 or 2017, I can& #39;t quite remember  https://abs.twimg.com/emoji/v2/... draggable="false" alt="🤣" title="Rolling on the floor laughing" aria-label="Emoji: Rolling on the floor laughing">)
https://abs.twimg.com/emoji/v2/... draggable="false" alt="🤣" title="Rolling on the floor laughing" aria-label="Emoji: Rolling on the floor laughing">)
And finally there& #39;s ion mobility spectrometry (IMS), a field that is so mind-bogglingly huge that I really need another #LipidTakeover to do it justice. But I will highlight a few select papers of the field that are pushing the boundaries in terms of lipid structure research.
A number of groups are using classic or drift time IMS to study lipids, including Groessl and colleagues who achieved near baseline separation of phosphatidylcholine regioisomers from porcine brain extract using high-resolution IMS
https://doi.org/10.1039/C5AN00838G">https://doi.org/10.1039/C...
https://doi.org/10.1039/C5AN00838G">https://doi.org/10.1039/C...
"Structures for Lossless Ion Manipulations" has also proven useful in separating both phospholipid C=C positional isomers and stereoisomers, a feat enabled by passing lipid ions through the equivalent of a 30.6 metre (!) drift tube length https://doi.org/10.3390/ijms18010183">https://doi.org/10.3390/i...
Differential ion mobility (DMS) is another IMS technique used to separate lipid isomers. Alex Shvartsburg of @WichitaState has used high-res DMS to separate lipid regioisomers, and both C=C positional and stereoisomers https://doi.org/10.1007/s13361-017-1675-2">https://doi.org/10.1007/s...
The Blanksby-Mitchell group have also tinkered with DMS and have used it to separate phospholipid regioisomers in natural extracts http://www.jlr.org/content/55/8/1668">https://www.jlr.org/content/5...
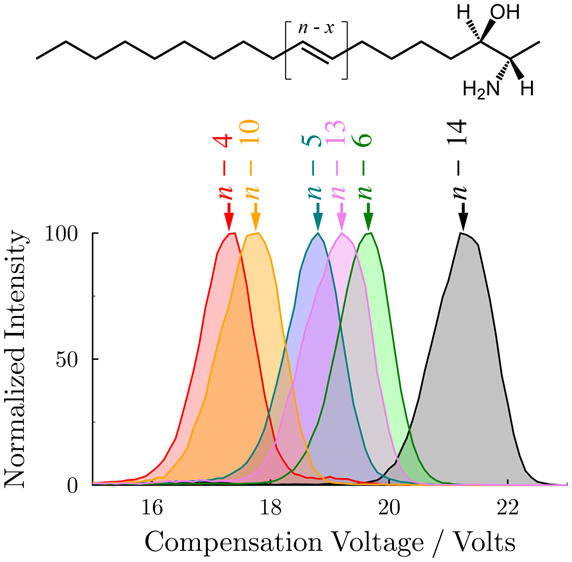
We& #39;ve also combined DMS with OzID to show that DMS can both separate 1-deoxysphingolipid C=C positional isomers and provide insights into their gas-phase structure (more from @bljpoad) https://doi.org/10.1021/acs.analchem.8b00469">https://doi.org/10.1021/a...
And last year we published work in @jlipidres showing the same usefulness of DMS for both C=C isomer separation and gas-phase structure analysis of very long to ultra-long chain neutral lipids found in meibum extracts. http://www.jlr.org/content/60/11/1968">https://www.jlr.org/content/6...
So that& #39;s it for this thread, and please do hit me up in the replies with any great new work in the field that I may have missed. In the next hour I will be awake, online, and ready to chat more about my experiences as an early-career researcher in Australia
 Read on Twitter
Read on Twitter