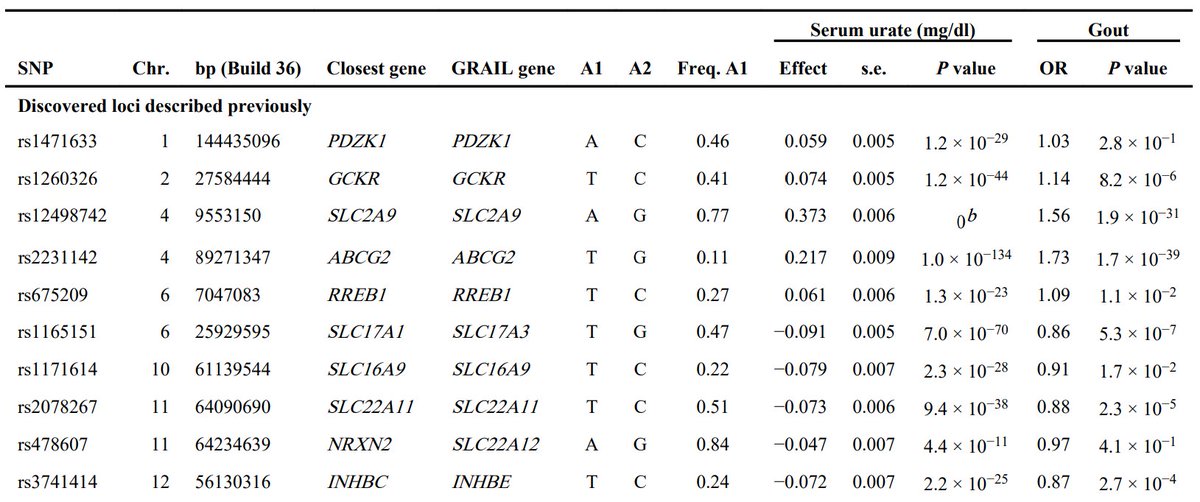
Something very comforting about the v high reliability and reproducibility of modern GWAS. I believe the first report of an association between SLC2A9 and uric acid and gout is this paper from a dozen years ago:
https://www.nature.com/articles/ng.106 ">https://www.nature.com/articles/... https://twitter.com/SbotGwa/status/1246783774018473990">https://twitter.com/SbotGwa/s...
https://www.nature.com/articles/ng.106 ">https://www.nature.com/articles/... https://twitter.com/SbotGwa/status/1246783774018473990">https://twitter.com/SbotGwa/s...
I guess that& #39;s the first to associate SLC2A9 with gout. @gwascatalog lists this as the first publication to link SLC2A9 to uric acid: http://www.ncbi.nlm.nih.gov/pubmed/17997608
The">https://www.ncbi.nlm.nih.gov/pubmed/17... GLUT9 gene is associated with serum uric acid levels in Sardinia and Chianti cohorts.
https://www.ebi.ac.uk/gwas/genes/SLC2A9">https://www.ebi.ac.uk/gwas/gene...
The">https://www.ncbi.nlm.nih.gov/pubmed/17... GLUT9 gene is associated with serum uric acid levels in Sardinia and Chianti cohorts.
https://www.ebi.ac.uk/gwas/genes/SLC2A9">https://www.ebi.ac.uk/gwas/gene...
In total, @GWASCatalog lists 79 studies with genome-wide significant hits for SLC2A9 and uric acid!
The strongest p-value reported in the gwas catalog is 1e-700, but this is incorrect. Yes, I get to mention my favorite pet peeve around underflow. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3663712/">https://www.ncbi.nlm.nih.gov/pmc/artic...
The strongest p-value reported in the gwas catalog is 1e-700, but this is incorrect. Yes, I get to mention my favorite pet peeve around underflow. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC3663712/">https://www.ncbi.nlm.nih.gov/pmc/artic...
Here& #39;s the table from the original paper. I calculate a p-value of about 8e-843. It would be nice to clean this up.
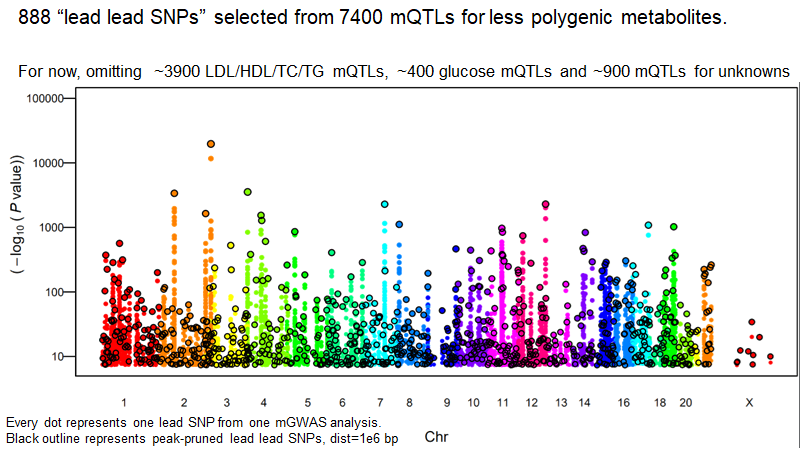
And even that is not the strongest p-value ever published for SLC2A9. That honor goes to Nasa Sinnott-Armstrong and team for the 38 biomarkers paper currently in biorxiv https://twitter.com/Eric_Fauman/status/1184814396087783427">https://twitter.com/Eric_Faum...
 Read on Twitter
Read on Twitter